- Home
- Services
- About
- News
- Contact
- Getting mtg arena on mac
- Whiteness of step up movies
- Mkvtoolnix download windows 7 32bit
- Does eastwest quantum leap work with reaper
- Word 2016 how to make a footnote
- Where can i download evernote for windows 10
- Intellij community edition download
- The recreated sinclair zx spectrum games
- Little girl in blue velvet 1978 movie
- Office 365 subscription prices
- Pajama sam 1 manual
- Star wars revisited adywan torrent
- Malwarebytes update problem 2020
- Double ended synthetic dreads red
- Hwo to set up out of office reply outlook 2013
- How can i upgrade my mac to sierra
- Download vmware workstation for windows vista
- What makes a serial killer snap
- Speed up clip in imovie for iphone
- Best youtube to mp3 converter program
- How to download house party custom stories
- Upgrade office 2007 32 bit to office 2010 64 bit
- I remember you keyshia cole mp3 download
- Best music mashups 2017
- Pokemon advanced adventure rom download mediafire
- Hevc codec for apple osx
- Convert 3 letter amino acid code to 1 letter


The other solution is to retrieve the mapping online (But the url and the html pattern may change through time): import reĭef three_to_one_online(three_letter_code): Ambiguous amino acids 3-letter 1-letter Amino acids included Codons included Any unknown Xaa X All NNN Asparagine or aspartic acid Asx B D N RAY Glutamine or glutamic acid Glx Z E Q SAR Leucine or isoleucine Xle J I L YTR ATH CTY coding codons can also be expressed by. Return mapping.upper() + three_letter_code.lower()] Conversion of three-letter aminoacid code into one-letter code and vice versa. The accepted codes in the input include the 20 standard amino acids and also B (Asparagine or Aspartic acid), converted to Asx, and Z (Glutamine or Glutamic acid) converted to Glx. The seq input length should be a factor of 3 or else resultsĭ = # you can add more Converts amino acid one-letter abbreviations to three-letter codes, e.g., L to Leu. The 3 letter code can be upper, lower, or any mix of cases '''Turn a three letter protein into a one letter protein. Here is a similar function to the one he provided that is all-inclusive and handles lower cases as well. Create a dictionary and use a for loop to read the string in blocks of three and translate. As far as your problem, Junuxx is correct.
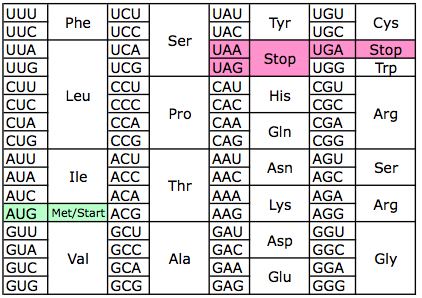
Unfortunately, there is no 'seq1' function (yet.) but I thought this might be helpful to you in the future. >'MetAlaIleValMetGlyArgTrpLysGlyAlaArgTer' I added this feature because the main program window only supports 3 letter amino acid abbreviations, but sequences obtained via web searches of sequence databases are typically in one letter notation. Example output in IDLE: >seq3("MAIVMGRWKGAR*") The Amino Acid Converter can be used to convert sequences of amino acid residues between 1 and 3 letter notation. Unfortunately, Biopython only has the function seq3(in package Bio::SeqUtils) which does the inverse of what you want. See Also AAString, GENETICCODE Examples See all the 3-letter codes AMINOACIDCODE Convert an AAString object to a vector of 3-letter amino acid codes aa <- AAString('LANDEECQW') AMINOACIDCODEstrsplit(as.character(aa), NULL. fasta file and then converting to one letter codes. Named character vector mapping single-letter amino acid representations to 3-letter amino acid representations. IN NO EVENT SHALL CHANG BIOSCIENCE OR ITS SPONSORS BE LIABLE FOR ANY DIRECT, INDIRECT, INCIDENTAL, SPECIAL, EXEMPLARY, OR CONSEQUENTIAL DAMAGES (INCLUDING, BUT NOT LIMITED TO, PROCUREMENT OF SUBSTITUTE GOODS OR SERVICES LOSS OF USE, DATA, OR PROFITS OR BUSINESS INTERRUPTION) HOWEVER CAUSED AND ON ANY THEORY OF LIABILITY, WHETHER IN CONTRACT, STRICT LIABILITY, OR TORT (INCLUDING NEGLIGENCE OR OTHERWISE) ARISING IN ANY WAY OUT OF THE USE OF THIS SOFTWARE, EVEN IF ADVISED OF THE POSSIBILITY OF SUCH DAMAGE.Ĭopyright © 2002-2004 Chang Bioscience, Inc.You may try looking into and installing Biopython since you are parsing a.
Convert 3 letter amino acid code to 1 letter software#
THIS SOFTWARE IS PROVIDED BY CHANG BIOSCIENCE ``AS IS'' AND ANY EXPRESS OR IMPLIED WARRANTIES, INCLUDING, BUT NOT LIMITED TO, THE IMPLIED WARRANTIES OF MERCHANTABILITY AND FITNESS FOR A PARTICULAR PURPOSE ARE DISCLAIMED. Thanks for using our software! Please let us know your suggestions and comments. Res Protein Letters 1.0 - Conversion between single and three-letter amino acid codes. Include all Primo 3.4, Abie 3.0, Heatmap Viewer, MicroHelper, Godlist Manager, label printing, and grade book. A great quick and practical reference for bench scientistsĪ collection of tools frequently used by bench biomedical scientists, ranging from centrifugationįorce conversion, molecular weight, OD, recipe calculators, to clinical calculators. The Electronic Protocol Book Table of contentsĪn electronic protocol book with 500 protocols andġ00 recipes. Mac: Internet Explorer 5.0, IE5.1.4 or higher (IE5.1 will not work because of a bug.) Can be used as content for research and analysis. PC: Internet Explorer 5.5 or higher and Netscape 4.08 or higher Collected from the entire web and summarized to include only the most important parts of it. Res Protein Letters 1.0: Converts Between Single- and Three-letter Codes Protein Sequences.Ĭlick here for more molecular biology tools.
- Home
- Services
- About
- News
- Contact
- Getting mtg arena on mac
- Whiteness of step up movies
- Mkvtoolnix download windows 7 32bit
- Does eastwest quantum leap work with reaper
- Word 2016 how to make a footnote
- Where can i download evernote for windows 10
- Intellij community edition download
- The recreated sinclair zx spectrum games
- Little girl in blue velvet 1978 movie
- Office 365 subscription prices
- Pajama sam 1 manual
- Star wars revisited adywan torrent
- Malwarebytes update problem 2020
- Double ended synthetic dreads red
- Hwo to set up out of office reply outlook 2013
- How can i upgrade my mac to sierra
- Download vmware workstation for windows vista
- What makes a serial killer snap
- Speed up clip in imovie for iphone
- Best youtube to mp3 converter program
- How to download house party custom stories
- Upgrade office 2007 32 bit to office 2010 64 bit
- I remember you keyshia cole mp3 download
- Best music mashups 2017
- Pokemon advanced adventure rom download mediafire
- Hevc codec for apple osx
- Convert 3 letter amino acid code to 1 letter
